OUR RESEARCH
Novel NADP redox sensors
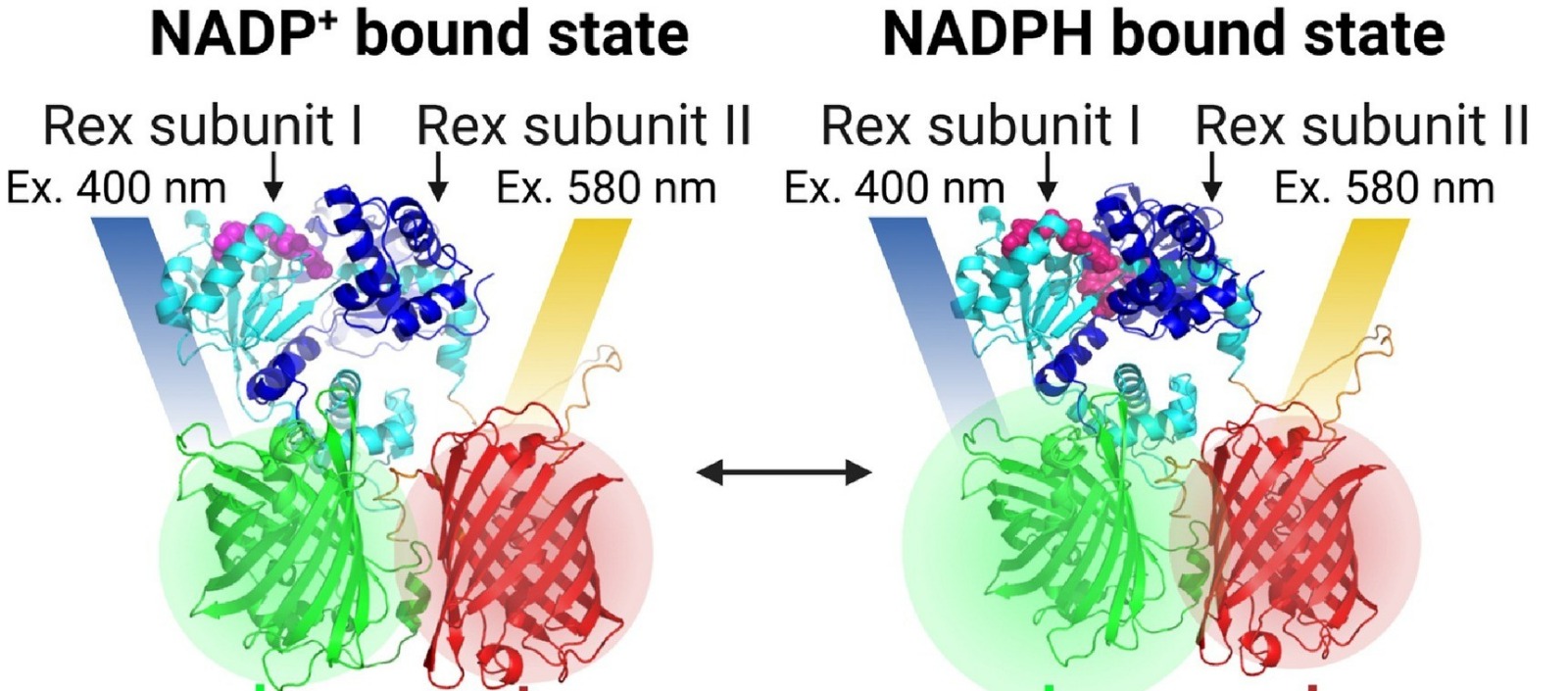
We developed the NAPstar family of genetically encoded fluorescent sensors that report NADPH/NADP⁺ ratios in real time. By turning fluorescence into a quantitative measure of cellular reducing power, we enable precise analysis of compartment-specific NADPH dynamics in living systems. We now work to develop novel NAPstars with improved sensitivity, compartment-specific targeting, and spectral variants suited for multiplexing.
Redox-dependent coupling of metabolism and cell division
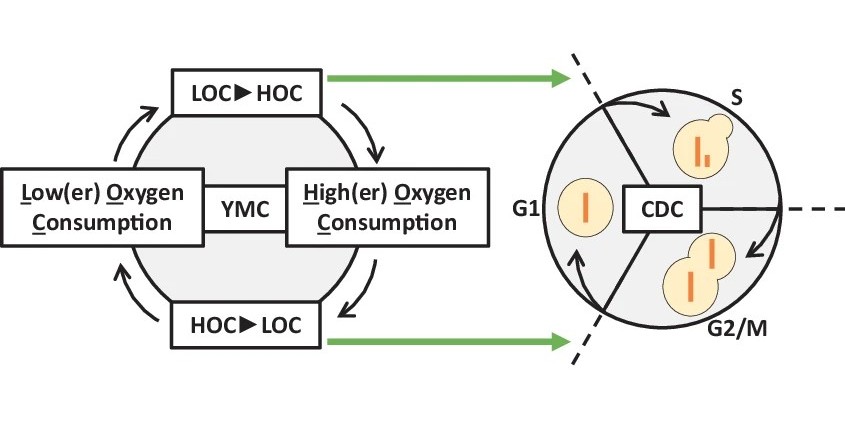
There is increasing evidence that self-sustained metabolic oscillations play an important role in regulating cell cycle entry and exit. We recently showed the peroxiredoxin-dependent redox signalling is important for coupling of cell division to metabolic oscillations. We are now working to understanding the mechanistic basis, with a focus on peroxiredoxin-dependent redox regulation of PKA.
Peroxiredoxin hetero-oligomerisation
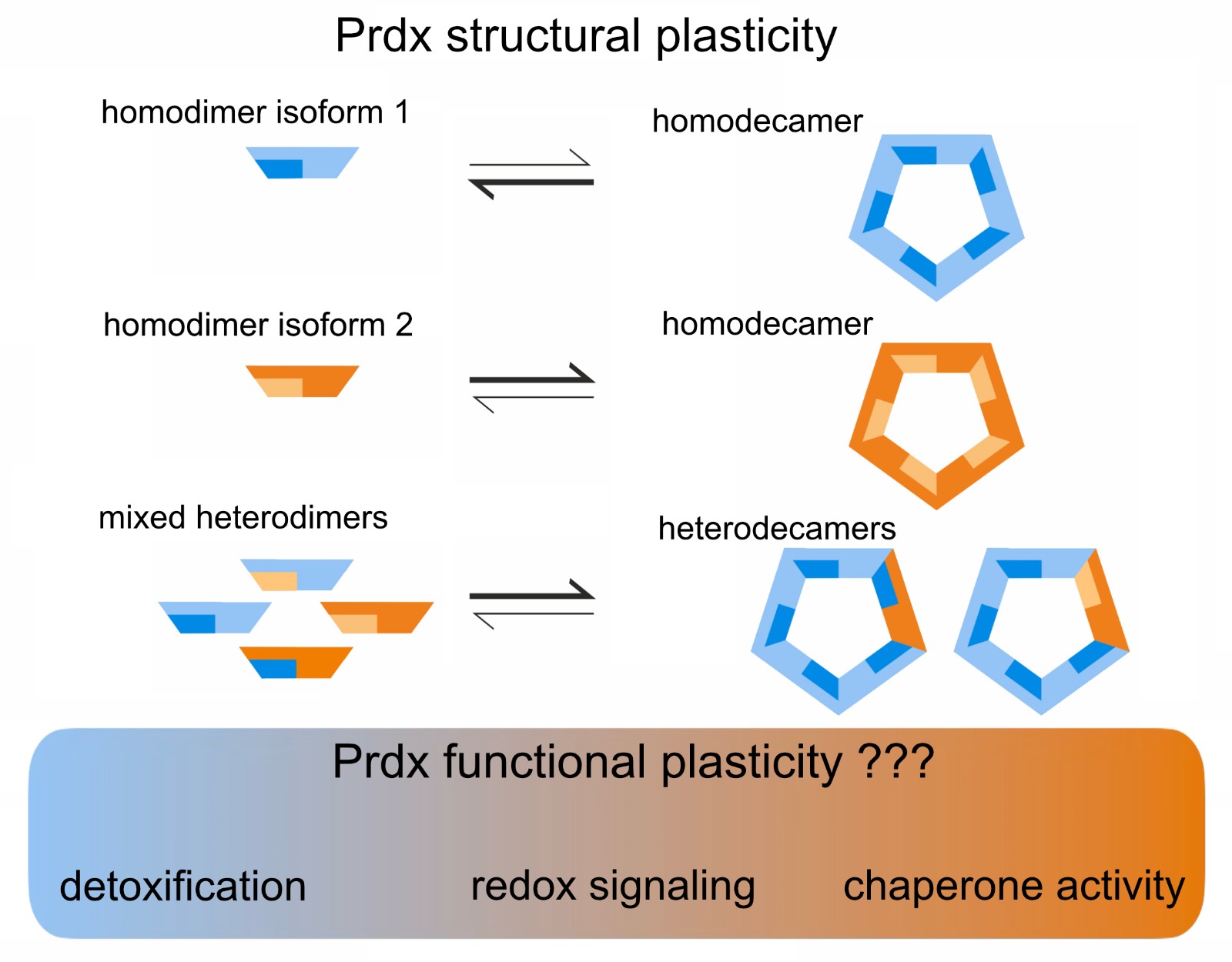
Peroxiredoxins are highly abundant peroxide-detoxifying enzymes with important roles in redox signaling and as chaperones. We recently discovered heterooligomerisation as a novel paradigm in eukaryotic peroxiredoxin biology. By combining structural biology, biochemistry and cell biology, we are investigating how oligomeric plasticity tunes peroxide detoxification, redox signalling and client protein interactions.
In vivo screening of enzyme structure–function
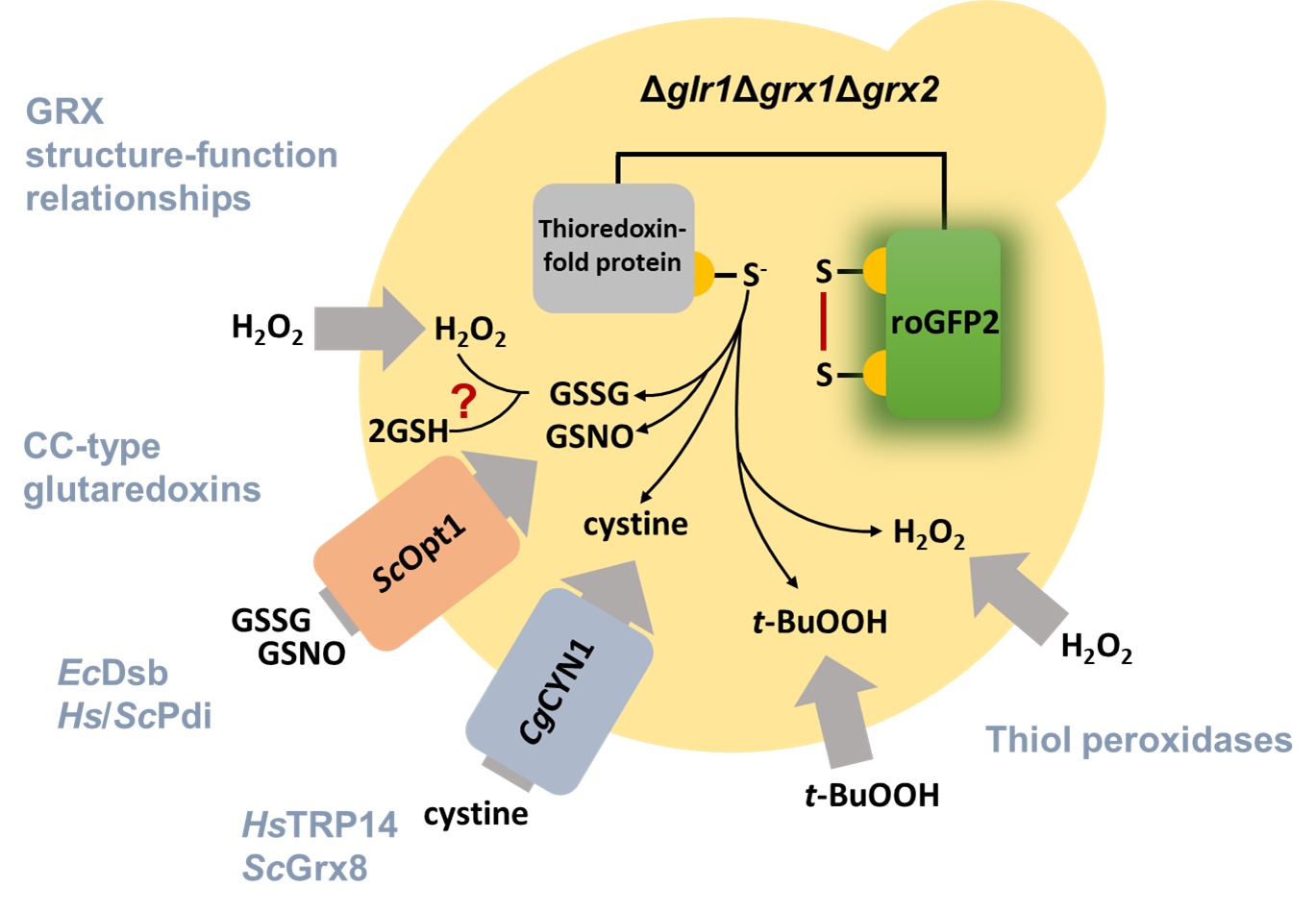
Our lab has developed yeast- and redox-sensitive fluorescent protein-based strategies to probe enzyme activity inside living cells. Using these methods in a screening approach we have been able to gain important new insights into structure-function relationships in diverse thioredoxin superfamily proteins. Future work will focus on the little explored role of post-translational modifications in the regulation of redox enzyme activity.
Red fluorescent redox-sensitive proteins
We are expanding the biosensor palette into the red and far-red spectral range, creating probes optimized for multiplexed experiments and deep-tissue imaging. These sensors overcome limitations of green-only probes, such as autofluorescence and shallow penetration, and are well-suited to studies in complex tissues and whole organisms. By engineering red fluorescent proteins to report on glutathione redox potential, H₂O₂, and NAD(P), we are developing powerful new tools for imaging redox processes alongside established green sensors, enabling multi-parameter readouts of cellular redox states.